Hello, I am trying to rotate a particle set by 120 degrees (2.0944 radians) around psi (Z-axis). I thought this was going to be straightforward but it’s not.
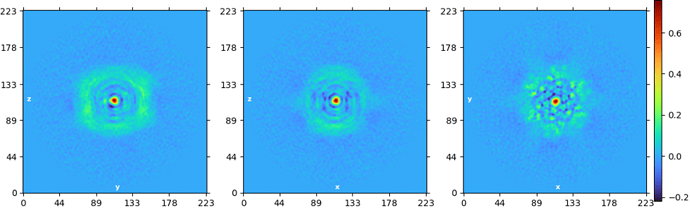
This is the original map:
I used the built-in interactive shell to load the particle cs file (as dataset_path) and then edit the euler angles using the following commands:
eulers = particle_dataset.data[‘alignments3D/pose’]
new_eulers = eulers
new_eulers[:,2] = eulers[:,2] + 2.0944
particle_dataset.data[‘alignments3D/pose’] = new_eulers
Then, I saved the new cs file:
particle_dataset.to_file(dataset_path)
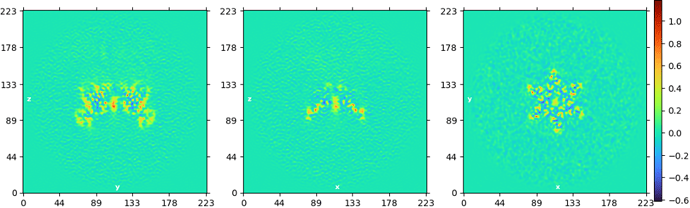
Then, I imported the new cs file and did a Homogeneous Reconstruction Only run. I expected that the map will now be rotated by 120 degrees around the Z-axis, but instead I get this:
Obviously, I’m missing something. Any guidance will be appreciated.
Owen
Hi @opornillos,
Great question. Since CryoSPARC stores orientations in axis-angle format, the correct transformation is a bit more complicated. You would have to take the alignments3D/pose, convert the axis-angles to rotation matrices, then compose that with a 120º rotation matrix around z – but there’s an easier alternative.
If you instead take the particles and pass them through a symmetry expansion job with C3 specified, then you will end up with a dataset of 3x the original size. Each particle will be rotated by 0º, 120º, and 240º, and each of these copies saved as a new particle. The sym_expand/idx field (with values 0, 1, or 2) will differentiate between the three. I don’t know the exact mapping between idx and the applied rotation angle, but idx of 0 should correspond to the 0º rotation, so you’re interested in the subset of the dataset with idx of either 1 or 2. You then have to subset the dataset by selecting only the particles with the desired idx: you can use the following snippet to do that:
from cryosparc_compute import dataset
Then load your symmetry expanded dataset (particle_dataset) and subset using:
new_dataset = particle_dataset.subset_query(lambda p : p['sym_expand/idx'] == 1) # or 2
Then save to disk and import the new cs file (can be done using Import Result Group).
Check out this thread for a similar request.
Best,
Michael
3 Likes
Hi @mmclean,
Thanks so much! This is exactly what I needed to do.
Best,
Owen