Okay, I’ve put together a script you can run to modify the results of the motion correction job and filter-out zero-sized files, here’s how to run it:
- Paste the following code into a text editor and make the noted subsitutions:
project_uid = '<PROJECT UID HERE>' datafile = '<OUTPUT DATA FILEPATH HERE>' passthrough_datafile = '<PASSTHROUGH OUTPUT DATAFILE HERE>' import os from shutil import copyfile from cryosparc_compute.dataset import Dataset project_dir = cli.get_project_dir_abs(project_uid) copyfile(datafile, f'{datafile}.bak') d1 = Dataset().from_file(datafile) keep_uids = set() for uid, path in zip(d1.data['uid'], d1.data['micrograph_blob/path']): full_path = os.path.join(project_dir, path) if os.path.exists(full_path) and os.stat(full_path).st_size > 0: keep_uids.add(uid) print(f'Found {len(keep_uids)}/{len(d1)} exposures to keep ({len(d1) - len(keep_uids)} empty files)') newd1 = d1.subset_query(lambda item: item['uid'] in keep_uids) newd1.to_file(datafile) copyfile(passthrough_datafile, f'{passthrough_datafile}.bak') d2 = Dataset().from_file(passthrough_datafile) newd2 = d2.subset_query(lambda item: item['uid'] in keep_uids) newd2.to_file(passthrough_datafile) print('Done filtering.')- Replace
<PROJECT UID HERE>with the ID of the project you are in, e.g.,P3 - Replace
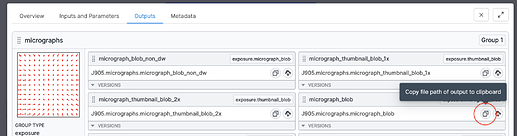
<OUTPUT DATA FILEPATH HERE>with the path to themicrograph_bloboutput of your motion correction job. Copy this value from the job’s “Outputs” tab, as indicated:
- Similarly replace
<PASSTHROUGH OUTPUT DATAFILE HERE>with the path from one of the outputs marked as “passthrough”, e.g., themscope_paramsoutput
- Replace
- Enter cryoSPARC’s interactive CLI with this command:
cryosparcm icli - Paste and enter the modified code from your text editor into the prompt
- Re-run the Patch CTF job
- Re-run the Template Picker job
Let me know if you have any trouble with that.