I’ve been testing a Live session on 5.0.1 and seem to have hit a bug. Using accepted exposures and particles from Live as input for the Junk Detector job, it failed at the end while compiling job outputs:
Timestamp: Feb 3, 2026, 11:12:25 AM
CryoSPARC Version: v5.0.1
P125 J17: Mic. Junk Detector
```
Traceback (most recent call last):
File "cli/run.py", line 112, in cli.run.run_job
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1581, in passthrough_outputs
job = passthrough_input(job, input_name)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1590, in passthrough_input
input_dset = load_input(job, input_name, slots="all")
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1739, in load_input
dsets = load_input_connections(job, input_name, slots=matching_slots, force_join=force_join)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1809, in load_input_connections
key: load_output(
^^^^^^^^^^^^
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1971, in load_output
metafile = find_output_result_metafile(job, output_name, result.name, version)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/bioxray/programs/cryosparc2/cryosparc2_worker/core/jobs.py", line 1871, in find_output_result_metafile
raise JobConnectionLoadError(
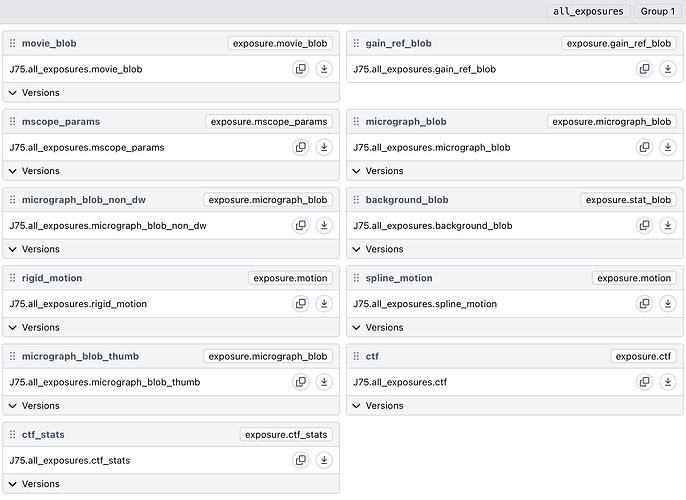
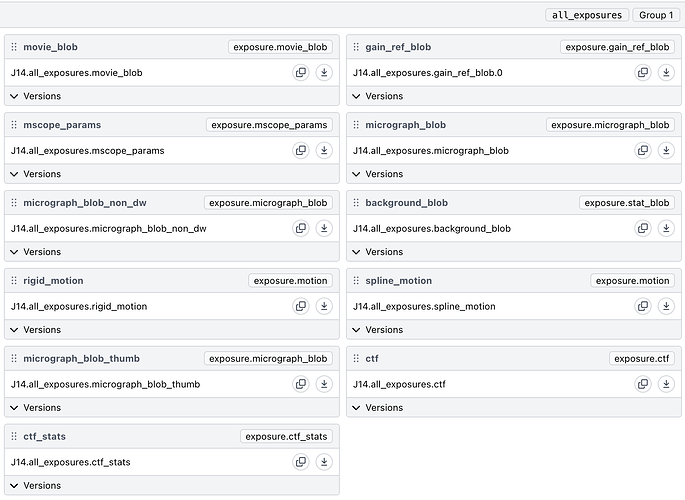
core.jobs.JobConnectionLoadError: Job P125-J15 (Live Exposure Export) Accepted Exposures output does not have result gain_ref_blob with version F
```
It refers to a missing gain_ref_blob, but the data was gain corrected on-the-fly and doesn’t (and shouldn’t) have a gain ref file in the Cryosparc project.
Marking the Junk Detector job as complete and using the output for Curate Exposures also produces an error before starting:
Timestamp: Feb 3, 2026, 11:41:37 AM
CryoSPARC Version: v5.0.1
P125 J18: Curate Exposures
```
Traceback (most recent call last):
File "cli/run.py", line 105, in cli.run.run_job
File "cli/run.py", line 210, in cli.run.run_job_function
File "/opt/cryosparc2/cryosparc2_master/compute/jobs/exposure_curation/run.py", line 68, in run
micrographs_dset = rc.load_input("exposures", required=True)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/run_context.py", line 321, in load_input
return jobs.load_input(
^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1739, in load_input
dsets = load_input_connections(job, input_name, slots=matching_slots, force_join=force_join)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1809, in load_input_connections
key: load_output(
^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1971, in load_output
metafile = find_output_result_metafile(job, output_name, result.name, version)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1871, in find_output_result_metafile
raise JobConnectionLoadError(
core.jobs.JobConnectionLoadError: Job P125-J17 (Micrograph Junk Detector) Labeled micrographs output does not have result micrograph_blob with version F
```
Curation of the exposures and particles directly from the Live exports start as expected, but also ends with an error while compiling outputs:
Timestamp: Feb 3, 2026, 11:48:52 AM
CryoSPARC Version: v5.0.1
P125 J19: Curate Exposures
```
Traceback (most recent call last):
File "cli/run.py", line 112, in cli.run.run_job
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1581, in passthrough_outputs
job = passthrough_input(job, input_name)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1590, in passthrough_input
input_dset = load_input(job, input_name, slots="all")
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1739, in load_input
dsets = load_input_connections(job, input_name, slots=matching_slots, force_join=force_join)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1809, in load_input_connections
key: load_output(
^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1971, in load_output
metafile = find_output_result_metafile(job, output_name, result.name, version)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/cryosparc2/cryosparc2_master/core/jobs.py", line 1871, in find_output_result_metafile
raise JobConnectionLoadError(
core.jobs.JobConnectionLoadError: Job P125-J15 (Live Exposure Export) Accepted Exposures output does not have result gain_ref_blob with version F
```
Anything I can do to continue processing on 5.0.1 or does it need to be fixed in a patch/update?
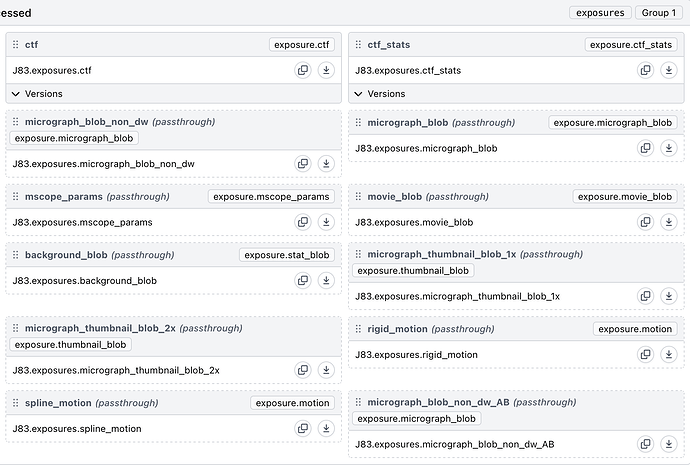
Update: it seems most/all jobs that work on the Live exposure exports fail - I’ve tried Exposure Sets, Curate Exposures, Manual Picker, Extract Particles, Blob Picker, Patch CTF Estimation and Micrograph Denoiser which all failed. The particles exported from Live seem okay and can be used for eg. 2D Classification and Ab-Initio jobs.
Update 2: Manual import of the same data does not lead to the same errors. After Patch Motion Correction and Patch CTF Estimation, downstream jobs seem to work as expected.