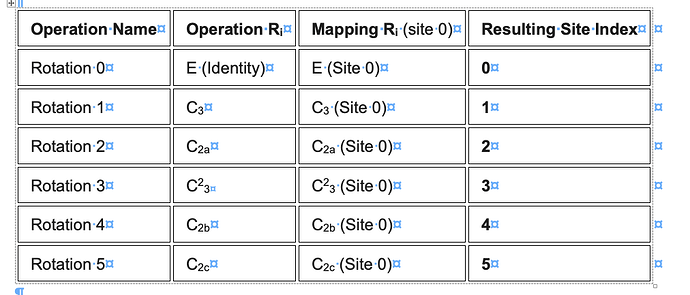
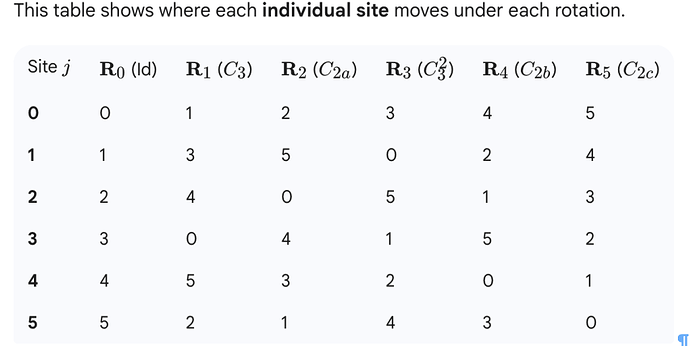
I have generated a symmetry-expanded set from a D3-symmetric protein complex. Then, after a series of classifications for a binding partner, I back-mapped the binding-partner-containing asymmetric units to the original structure to define a set of 16 structurally distinct D3 states.
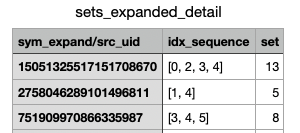
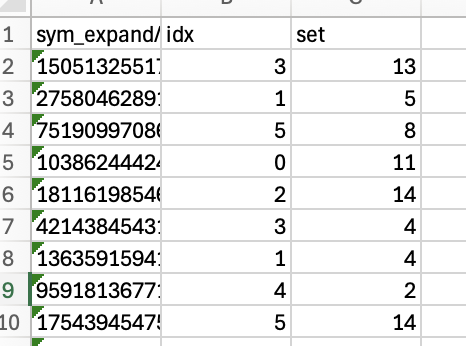
Currently, I have lists of UIDs and specific idx values for the symmetry expanded particles in a .csv format. I would like to use the original symmetry expanded particle set to make a cryoSPARC-ready .cs file that contains only the particles with the UIDs and idx values listed in my .csv file—i.e., copy and paste the particle information for the subset of particles listed in the .csv file into a new .cs file so that I can then upload it into Cryosparc for further processing.
I am assuming Cryosparc tools would be the most direct way to do this, but I am not sure how to implement it. What would be the best way to do this?
Thank you!