Hi everyone,
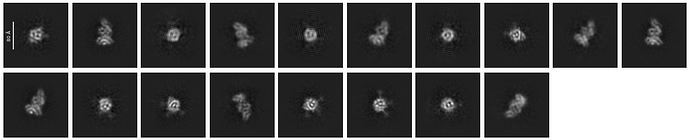
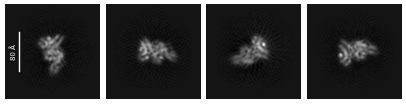
I am processing a small protein, about 61 kDa. I have a reasonably good number of particles and some promising 2D classes, but I have noticed that some of the apparent top/bottom views pick up a fair amount of noise (attached), even after tightening the circular mask parameters.
So far I have tried adjusting the number of iterations, force max over poses/shifts, clamp solvent, and enforce negativity, using a bigger box six=ze based on some recommendations I have come across in this forum. My ab initio classes actually look fairly decent. I followed the HR-HAIR workflow from Oliver Clarke | bioRxiv included those results below as well. I tested runs with about 5–8 classes.
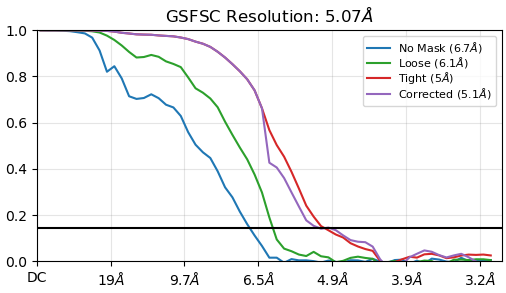
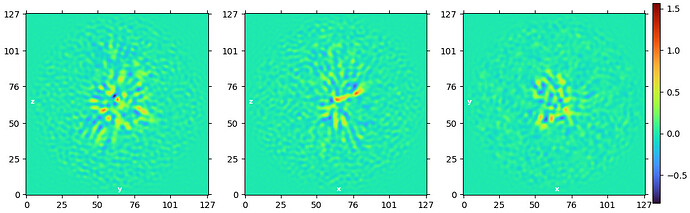
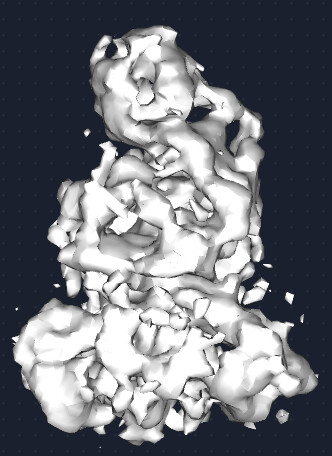
Where things start to fall apart is after that step. My downstream 3D refinements become very noisy, look overfit, and develop a streaky appearance. I have tried Homo Refinement settings(results attached below), described in the same paper, as well as Hetero Refinement against junk classes, and I also tested a soft static mask, and NU-refinement.
At this point, I honestly feel like the ab initio models look better than the later refinement jobs, which makes me think I may be pushing the data in the wrong direction after the initial reconstruction stage.
I have learned a lot from reading through this forum, and many of the tips here have already helped me get this far. I would really appreciate any advice on what to try next, especially for improving refinement quality and avoiding the noisy/streaky reconstructions. I am happy to provide more details about my workflow, parameters, or intermediate results if that would help!