Hi all,
Could someone please help us to figure out if our structure has an orientation issue? I attached the orientation distribution diagram here. More importantly, are there any widely accepted standard critera to judge if a map has orientation issue other than just looking at the map?
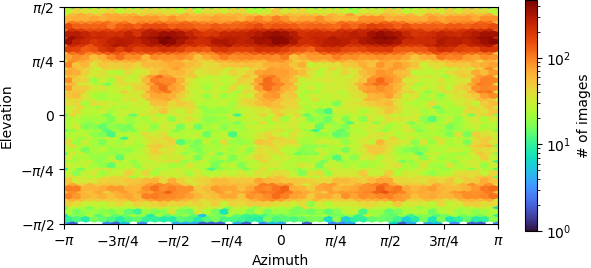
Thank you very much.
1 Like
Hi Zhengshan,
This doesn’t look too bad to me. Why do you suspect an orientation issue? Given it looks like your protein has fourfold (?) symmetry, if you have poor/anisotropic density I would first consider pseudosymmetry or interdomain flexibility as possible explanations.
To your second question, I would suggest looking up Dmitry Lyumkis’s work (3D-FSC and related topics). I wouldn’t say there is a standard as such, but there are certainly some useful tools for assessment of anisotropy.
Cheers
Oli
2 Likes
To add to Oli’s advice, you may also wish to read up on Naydenova and Russo’s cryoEF, which complements Dmitry’s informative 3D-FSC analysis. Anecdotally, we’ve seen a practical “resolvable” limit of E~0.7, although this will depend on the size/symmetry/flexibility of the particle.
(cryoEF extracts angles from .star files, so there will be some pyem conversion required.)
Cheers,
Yang
3 Likes
Hello, if your protein is not too small (~200KD), you can solve the data loss problem caused by the orientation advantage by tilting the sample stage, but this still cannot retrieve all the data, and the results will still be wrong Perfect, our research group has developed an algorithm that can restore missing data and make the reconstruction result better. If possible, we can cooperate. Contact information: caoweili20@mails.ucas.ac.cn
Hi All,
Can I ask how to convert the orientation star file from a Relion Refine3D to the input for cryoEF? Thanks in advance!
Hi,
Katerina has included scripts, e.g. extractAngles.sh, that you’ll find in the distribution package. I would suggest reading the manual for exact usage.
Cheers,
Yang
Yang,
Thank you very much!!!
I did not pay enough attention although I downloaded the package.
