Hi,
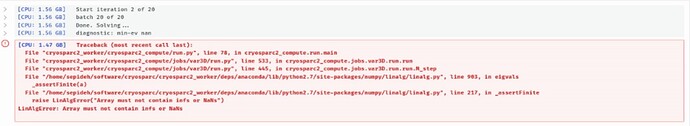
I am getting the following error while running 3D variability:
[CPU: 7.63 GB] Traceback (most recent call last):
File "cryosparc2_worker/cryosparc2_compute/run.py", line 85, in cryosparc2_compute.run.main
File "cryosparc2_worker/cryosparc2_compute/jobs/var3D/run.py", line 533, in cryosparc2_compute.jobs.var3D.run.run
File "cryosparc2_worker/cryosparc2_compute/jobs/var3D/run.py", line 445, in cryosparc2_compute.jobs.var3D.run.run.M_step
File "/users/svc_cryosparc/software/regular/cryosparc2_worker/deps/anaconda/lib/python2.7/site-packages/numpy/linalg/linalg.py", line 1058, in eigvals
_assertFinite(a)
File "/users/svc_cryosparc/software/regular/cryosparc2_worker/deps/anaconda/lib/python2.7/site-packages/numpy/linalg/linalg.py", line 218, in _assertFinite
raise LinAlgError("Array must not contain infs or NaNs")
LinAlgError: Array must not contain infs or NaNs
It happens in somewhat random iteration have seen it happening in iteration 5, 6 and 8 in different trials. I tried different queues.
The dataset however behaves fine and refines nicely in 3D refinement. Any idea?
Best,
David