Hi all,
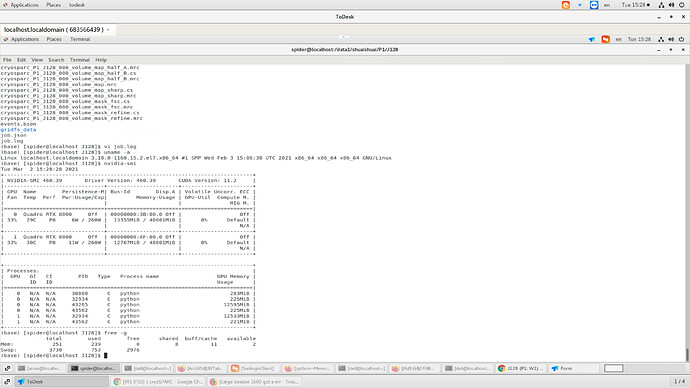
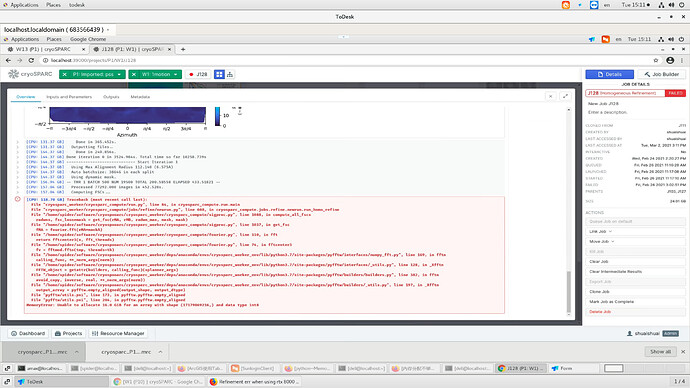
When I using cryosparc to do refinement with boxsize 1600,I got a err like blew.My Gpu is rtx8000,and my memory is 256 like your recommend in website,could give me some advice howe to fix that?
thanks very much.
[CPU: 102.46 GB] Traceback (most recent call last):
File "cryosparc_worker/cryosparc_compute/run.py", line 84, in cryosparc_compute.run.main
File "cryosparc_worker/cryosparc_compute/jobs/refine/newrun.py", line 327, in cryosparc_compute.jobs.refine.newrun.run_homo_refine
File "/home/spider/software/cryospooarc/cryosparc_worker/cryosparc_compute/newfourier.py", line 416, in resample_resize_real
return ZT( ifft( ZT(fft(x, stack=stack), N_resample, stack=stack), stack=stack), M, stack=stack), psize_final
File "/home/spider/software/cryospooarc/cryosparc_worker/cryosparc_compute/newfourier.py", line 119, in ifft
return ifftcenter3(X, fft_threads)
File "/home/spider/software/cryospooarc/cryosparc_worker/cryosparc_compute/newfourier.py", line 92, in ifftcenter3
v = fftmod.irfftn(tmp, threads=th)
File "/home/spider/software/cryospooarc/cryosparc_worker/deps/anaconda/envs/cryosparc_worker_env/lib/python3.7/site-packages/pyfftw/interfaces/numpy_fft.py", line 295, in irfftn
calling_func, **_norm_args(norm))
File "/home/spider/software/cryospooarc/cryosparc_worker/deps/anaconda/envs/cryosparc_worker_env/lib/python3.7/site-packages/pyfftw/interfaces/_utils.py", line 128, in _Xfftn
FFTW_object = getattr(builders, calling_func)(*planner_args)
File "/home/spider/software/cryospooarc/cryosparc_worker/deps/anaconda/envs/cryosparc_worker_env/lib/python3.7/site-packages/pyfftw/builders/builders.py", line 545, in irfftn
avoid_copy, inverse, real, **_norm_args(norm))
File "/home/spider/software/cryospooarc/cryosparc_worker/deps/anaconda/envs/cryosparc_worker_env/lib/python3.7/site-packages/pyfftw/builders/_utils.py", line 261, in _Xfftn
flags, threads, normalise_idft=normalise_idft, ortho=ortho)
File "pyfftw/pyfftw.pyx", line 1223, in pyfftw.pyfftw.FFTW.__cinit__
ValueError: ('Strides of the output array must be less than ', '2147483647')