Dear community!
I recently posted an topic about recentering 2D classes; the method I tried seemed good enough then, but it’s not quite good enough, we still want to find a way to manually recenter the classes.
I recenter each class by running the following python script
import numpy as np
import sys
def load_cs(cs_path):
print(f"[INFO] Loading CS file: {cs_path}")
arr = np.load(cs_path)
print(f"[INFO] Loaded array with {arr.shape[0]} particles and dtype fields:")
print(arr.dtype)
return arr
def save_cs(out_path, arr):
print(f"[INFO] Saving modified CS file to: {out_path}")
np.save(out_path, arr)
print("[INFO] Save successful.")
def shift_one_class(particles, target_class, add_shift):
if "alignments2D/class" not in particles.dtype.fields:
raise ValueError("[ERROR] No 'alignments2D/class' field — this is not 2D classification output.")
if "alignments2D/shift" not in particles.dtype.fields:
raise ValueError("[ERROR] No 'alignments2D/shift' field found.")
updated = particles.copy()
count_modified = 0
for i in range(len(particles)):
cls = particles["alignments2D/class"][i]
if cls == target_class:
old_shift = particles["alignments2D/shift"][i]
new_shift = old_shift + add_shift
updated["alignments2D/shift"][i] = new_shift
count_modified += 1
if i % 50000 == 0:
print(f"[LOG] Processed {i} particles...")
print(f"[INFO] Done shifting class {target_class}.")
print(f"[INFO] Modified {count_modified} particles.")
return updated
def main():
if len(sys.argv) < 6:
print("Usage:")
print(" python shift_one_class.py input.cs output.cs class_id dy dx")
print("Example:")
print(" python shift_one_class.py particles.cs out.cs 0 -50 0")
return
in_cs = sys.argv[1]
out_cs = sys.argv[2]
target_class = int(sys.argv[3])
dy = float(sys.argv[4])
dx = float(sys.argv[5])
add_shift = np.array([dy, dx], dtype=np.float32)
print(f"[INFO] Target class: {target_class}, shift = (dy={dy}, dx={dx})")
particles = load_cs(in_cs)
updated = shift_one_class(particles, target_class, add_shift)
save_cs(out_cs, updated)
if __name__ == "__main__":
main()
And created another csg file to import back to cryosparc
created: 2025-11-26 08:58:37.067915
group:
description: All particles that were processed, including alignments across all
classes.
name: particles
title: All particles
type: particle
results:
alignments2D:
metafile: '>shifted-particles.cs'
num_items: 35395
type: particle.alignments2D
blob:
metafile: '>shifted-particles.cs'
num_items: 35395
type: particle.blob
ctf:
metafile: '>shifted-particles.cs'
num_items: 35395
type: particle.ctf
location:
metafile: '>J226_passthrough_particles.cs'
num_items: 35395
type: particle.location
ml_properties:
metafile: '>J226_passthrough_particles.cs'
num_items: 35395
type: particle.ml_properties
pick_stats:
metafile: '>J226_passthrough_particles.cs'
num_items: 35395
type: particle.pick_stats
version: v4.7.1
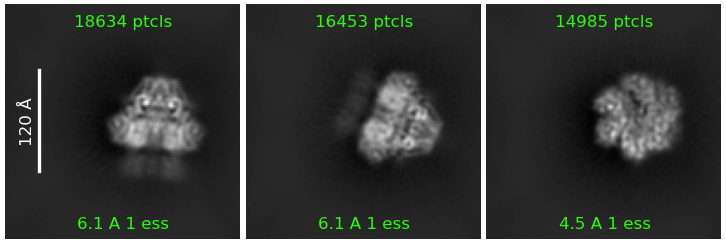
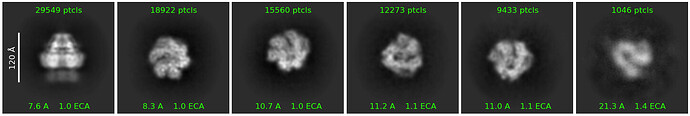
However, when I ran a 2D classification to see the changes, every class still landed dead-center, even though the shift I applied should have moved it outside the box.
How do I manually recenter the 2D classes?
Any suggestions would be great!