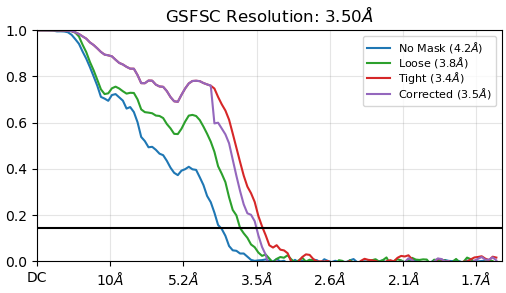
Hello, everyone. During data processing, I found that proteins offer orientation advantages. The Nu-refinement resolution is 3.5A. However, this orientation advantage problem needs to be solved. Do you have any good suggestions to solve this problem? Thank you very much!
this “advantage” which we often call bias can be solved in 2D class, simply identify which view is most prominent and do not select it for next steps. Also can be done automatically with “rebalance orientations”. Also can be fixed at sample prep by including detergents/amphipols/LEA/salmon sperm DNA etc.
my favorite trick requires a few steps: 2D select a lot of the dominant view - run ab initio. 2D select a smaller subset of particles that you think has ~equal of all views - run ab initio. run ab initio again but only “use this many particles” 10. Provide model 1, model 2, and a few copies of model 3 as total junk to a heterogeneous refinement job. the het refine will automatically build a preferred orientation class which will help to remove that bias, and will also provide an isotropic map of all views, and will also sort out junk/non-particles. This doesn’t always work on the first try, but if you keep trying to provide biased, isotropic, junk, high res, low res REFERENCES to the job, then you can get a good result and iterate over and over. I do this during data collection to get a good cleaning system, then apply it to the full particle set at the end of collection and get the same results.
Thank you for your suggestion, I will handle it according to the method you said, and I have some good news to share with you.