Hi all,
I am having issues with classification and refinement of a homodimer as it seems to want to align much more strongly on one of the monomers.
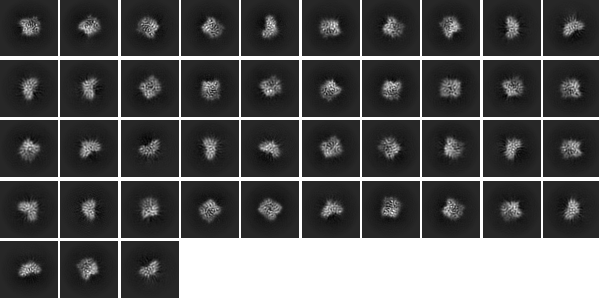
When I try to take these classes for use in heterogeneous or homogeneous refinement I end up seeing the same issue with the volumes that are spit out and quite a bumpy FSC curve. I believe we definitely need to collect more data for this project, but at this stage I’d like to know if anyone has some suggestions regarding parameters that could be tweaked to try and obtain more even alignment.
Cheers,
SW
Hi Sean,
If it is a ~C2 symmetric homodimer and the size of the monomer is sufficient (~>=50kDa), you might try symmetry expansion followed by local refinement in C1 with a mask covering only one of the monomers.
Cheers
Oli
1 Like
Thanks for the suggestion! I will give it a shot.
I’d also suggest if you are using defaults with Class2D, try a run marginalizing over poses and shifts (Force/Max off) with a larger number of iterations (40) and a larger batch size (200-400). Often gives better results for small, low SNR particles
Oli
1 Like
Hi Oli,
I’ve only ever worked on fairly small things so came across your twitter post with those parameters a while back. Have been a game changer! The classes above do not marginalize over poses, but I used 40 iterations and 500 batch size.
Thanks!
SW
I would also suggest restraining the alignment resolution to 8 - 12 A.
1 Like
Hi Oli,
I exported the particles I wanted to use and used pyem to generate a star file, used relion_particle_symmetry_expand to make a new star file, and then tried to import back into cryosparc, but it’s now not recognizing the file paths in the file. Do I need to edit the star file to contain absolute paths for the images rather than the “>Jxxx/…/xxx.mrc” that it currently has?
Cheers,
SW
Either that, or move the star file to where those relative paths make sense before importing.
Alternatively, you can just do the symmetry expansion in cryosparc if you are using the latest version?
Oli
Oops! I had just googled “symmetry expansion cryosparc” and only came up with threads where you were asking for it as a feature! I’ve done it now, but unfortunately I’m still in a position where I’m having a hard time determining what the boundary of the dimer is in 3D in order to mask accurately. I’m trying to do some homology modelling to have something to guide me.
hmm - you might try using the 3D variability analysis? This can be helpful for identifying protomer boundaries
It’s also OK to have some overlap, even preferable because you might discover some heterogeneity at the interface.
1 Like
Thank you so much for the suggestions! I’ve primarily done crystallography in the past, so I’m still very much just learning how to process EM data.
