Hi,
I was wondering if there is an easy way to select all particles assigned to a certain class of a 3D ab initio run to carry out further classification of those particles with another ab initio run? I guess you can just play around with the class probability threshold, but I was wondering if there is an easy way of just selecting all the particles into a specific class.
Thanks a lot in advance
Best wishes,
Rafa
Hey Rafa,
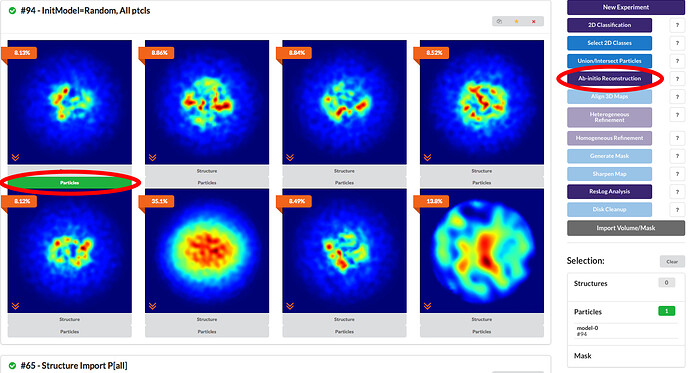
You would click “Particles” under the desired 3D class that you want to do further work on. Then you click on “Ab-Initio Reconstruction” to execute further classification on this particle subset.
Sincerely,
Nicholas Gao
Hi Nicholas,
Thanks a lot for your reply! However, if you do that, when later you run the next ab-initio job, it will only select the particles assigned to that class with a probability higher than what you specify in the class probability threshold parameter of the ab-initio job (by default 0.9, which in my case I have seen can leave out quite a few particles).
What surprises me is I have found cases where after running an ab-initio with 5 classes for example, including all the particles with a class probability threshold of 0.2 for one of the models does not include all the particles! This is unexpected, as as far as I understand it implies for those particles assigned to that specific class, they class for which they have the highest probability value is another class
Best wishes,
Rafa
it implies for those particles assigned to that specific class, they class for which they have the highest probability value is another class
It implies their maximum probability is that class, but that the membership probability is less than 0.2. If you really want all the particles, you can use a threshold of 0.
Interesting! However, I have found that by setting that parameter to 0, all the particle from all the classes are taken for the next round of classification! Instead of all the particles of a single, selected class… Any idea what could be going on here?
Thanks a lot in advance!
I have found that by setting that parameter to 0, all the particle from all the classes are taken for the next round of classification!
Oh, my apologies then. What I described is how I understood Ali’s intent…it’s also what my csparc2star.py program does when converting class assignments for Relion.
There are two alternate, conceptual operations:
- All particles with at least threshold probability of class membership
- Only particles with threshold probability and maximum probability for the class
Maybe @apunjani can comment as to his actual intent here.
I also interpreted it in the same way than you at the beginning! But then found that setting it to 0 led to selecting all the particles. It would be great if @apunjani can comment on this! Since with the other interpretation you would expect that when classifying a set of particles into say 5 models, by using a threshold of 0.2 you would be selecting all of the particles assigned to that class. Otherwise, if there are particles assigned into this class with a probability under 0.2, the probability for at least one other class will be necessarily higher than 0.2 (since all the probabilities should add up to 1 if I understand correctly)!
Best wishes,
Rafa
Hi @Rafael-Ayala, @DanielAsarnow,
Currently when you select “Particles” from the experiments page as input for a further ab-initio experiment, the new experiment will select a particle as input, if the sum of probabilities for that particle across all selected classes (if more than one class is selected) is greater than the threshold parameter in the new experiment (by default 0.9). This corresponds to #1 in Daniel’s enumeration.
If you set the threshold at 0, you will get all the particles from all classes. Setting it at 0.2 and selecting one of five classes will give all particles that have greater than 20% confidence of being in that class (even if they have maximum probability in a different class). The class assignment probabilities do sum to one.
Maybe what we can do is add an option alongside the probability threshold to set whether to only keep particles that have maximum probability in the selected class(es). This way both Daniel’s #1 and #2 can be used, if desired.
Ali
Hi @apunjani
Thanks for your reply! That would be a very handy option !
Best wishes,
Rafa