Hi all
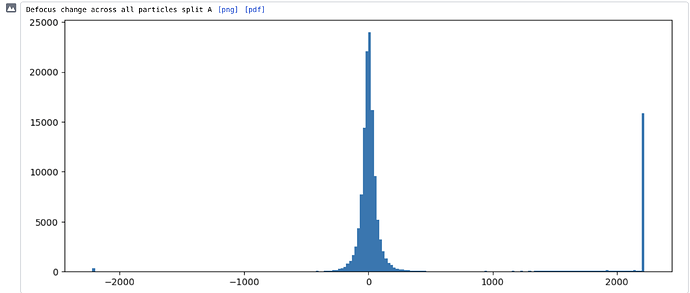
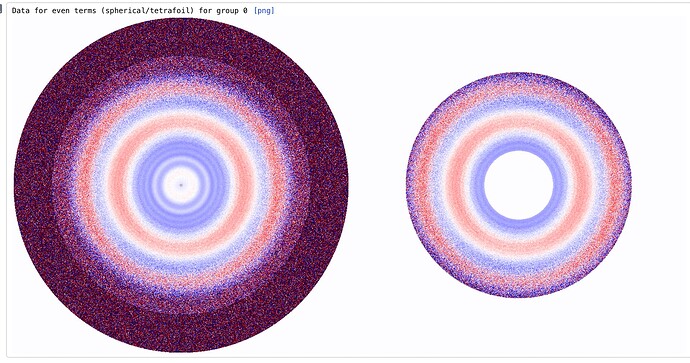
I have a misbehaving local CTF refinement job - most of the particles have defoci that refine to sensible values, but some are way off (see attached). I then see uncorrectable weirdness (almost like Thon rings) in the higher order aberration plots, which I think may be due to severely incorrect defocus estimates for this subset of particles.
I suspect that these particles originate from micrographs where CTF estimation has failed in some way (playing around with the usual parameters in Curate Micrographs did not do the trick, so I suspect it is something non-obvious). I would like to select these particles so I can either identify where they came from, fix the problem, or remove them.
Currently I don’t think there is a way to do this, as the change in defocus is not recorded as a parameter. Would this be possible to add for a future release, perhaps coupled to a mechanism to inspect and select particles based on metadata (of which this is one example)?
Cheers
Oli
1 Like
Hi Oli,
Conceptually, as a workaround, could you compare particle data from iteration n and iteration (n-1), specifically the defocus fields, and identify those for which the modulus of the difference is greater than e.g. 1000? For .star files, I imagine you could script it with awk, but I’m not familiar with how to work with .cs files.
Cheers,
Yang
1 Like
Yes - it should be possible by awking the particle.star, it’s on the to-do list… or maybe there is a way using cryosparc-tools…
Hi all,
It should be possible to do this via cryosparc-tools. At a high level, one would need to:
- Load the input and output particle stacks (to the local CTF Refinement job) into a python script, ensuring that the
ctf fields are loaded in
- Get the defocus parameters
ctf/df1_A and ctf/df2_A for input and output stacks
- Calculate the per-particle defocus as the mean of these two (i.e.
defocus_in = (particles_in['ctf/df1_A'] + particles_in['ctf/df2_A'])/2) for input and output stacks. (This mean is between the defocus along the major and minor axes per-particle).
- Calculate the difference in this quantity between input and output stacks (
defocus_change = abs(defocus_in - defocus_out))
- Using logical operations (e.g.
NumPy masking) on the output particle stack to remove the particles indices associated with the entries of the defocus_change array that exceed some threshold, e.g. 1000Å)
- Output the resulting subsetted particle stack through a CryoSPARC Tools External Job to a CryoSPARC Workspace
Best,
Michael
1 Like
Thanks @mmclean! Trying this now, running into some presumably newbie issues: Error loading particles using cryosparc-tools
What I would like to do is to have particle outputs created - one with the remaining subset of particles after outlier removal, and one with the outliers themselves. That way I can then use Inspect Picks to see why they are causing trouble
Cheers
Oli
Ended up doing this using awk as per @leetleyang’s suggestion:
After converting the particle.cs files of the iteration before and after local defocus refinement to star using pyem:
paste J649_006_particles.star J649_007_particles.star | awk 'function abs(v) {return v < 0 ? -v : v} {diff = abs($7 - $(NF/2+7)); if (diff < 1000) print $0}' > output_1000.star
This outputs all the particles with <1000Å absolute change in defocusU (which in this case is field #7) from the first iteration star. Don’t need to average them together, as local CTF does not refine astigmatism. Can be easily modified to just output the ones with greater than a certain defocus difference. After generating the star, you need to replace the header with that from the first file, as it gets a bit messed up.
Cheers
Oli
(PS: I needed to use the --inverty flag in csparc2star.py in order for the particle locations to match the micrographs upon re-import)
1 Like