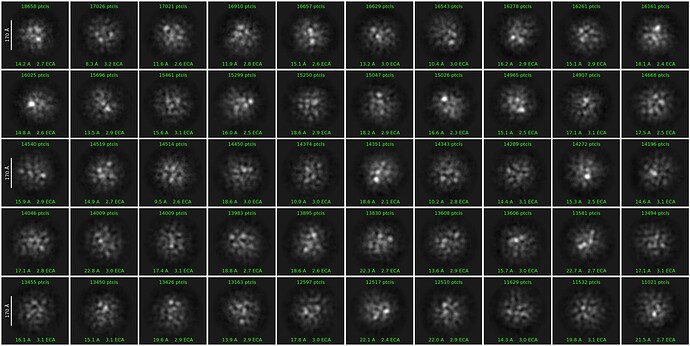
Hello,everyone, I am processing a Cryo-EM dataset of a protein complex around 400Kd, even though the micrograph is not good enough, the result after 2D classification is much worse than I expect. Does anyone have experience handling similar data? Where can we make improvements?
Many thanks in adavance!
show a micrograph and a picked micrograph. this could be anything… box size incorrect? if you increase batch size to 1000 do you get 1 good class? did you template pick (or just blob)? did you use thresholds to limit the amount of particle picks to those which you expect are real (can be extra conservative at first-pass to get good templates for future picking). What was wrong with the micrographs? 2D should be a very low bar, easy to get good classes for even bad ice, heterogeneous data, small or large particles. This suggests bad picking or incorrect parameters for extraction
excellent points by @CryoEM2
did you inspect the pics or put the pics + micrographs in manual picker ?
if so are a lot on the carbon ? it seems like these 2D classes resemble picks on carbon to me.
It is hard to miss with blob picker if you know about the size say for 400 kDa, e.g. 150-300 A diameter => extract 600pix with Fourier crop to 100 => run it through 2D.