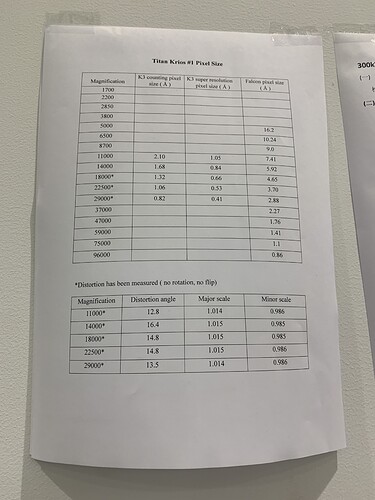
For our K3 data, we need to do the Distortion correction in the motioncor2 to obtain the right astigmatism evaluation.
But I don’t know how to do it in cryoSPARC.
Motioncor2 has the -Mag option to correct the Distortion.
-Mag
- Correct anisotropic magnification by stretching
image along the major axis, the axis where the
lower magificantion is detected.
- Three inputs are needed including magnifications
along major and minor axes and the angle of the
major axis relative to the image x-axis in degree.
- By default no correction is performed.
One option is to have motioncor2 output an aligned stack. If you just want to use alignparts_lmbfgs, or there is some other reason not to import motioncor2 aligned sums. You could also use mag_distortion_correct on your movies directly.
We tried the motioncor2’s distortion in Relion. But we obtained a bad result after polish in Relion.
I don’t know if the local motion correction could benefit this in cryoSPARC.
Hi Fengjiang,
That’s likely because motioncor2 doesn’t alter the stacks, and polishing doesn’t have any way of knowing about mag anisotropy. If you generate a gain and distortion corrected stack using mag_distortion_correct as @DanielAsarnow suggested, then that should work with either polishing in Relion or local motion correction in cryoSPARC. Only downside is you will need a lot of space to store the stacks, assuming you were previously storing non gain corrected and TIFF compressed stacks.
Cheers
Oli
Hi Oli,
That’s sounds great! But I didn’t find this mag_distortion_correct option.
Could you tell me where is it?
it’s not a cryosparc option - it is a separate package by Niko Grigorieff and co (http://grigoriefflab.janelia.org/magdistortion). I believe it may also be part of cistem now but I haven’t checked that. Good luck!
Cheers
Oli
@DanielAsarnow @olibclarke Thanks a lot! I going to try it.
Hi all - all the various aberration correction tools are in the development pipeline for cryoSPARC, including anisotropic mag correction (which will, like Relion, be done as part of refinement rather than to the micrographs themselves). For now though the workflow you described is likely the best approach.