Dear CryoSPARC team,
I am currently trying a non-standard workflow. Specifically, I converted particle-subtracted particles mrc stacks into standalone .mrc micrographs and attempted to import them into CryoSPARC as micrographs.
So far:
-
Working jobs: Import Micrographs, Topaz Denoise, Patch CTF Estimation, Blob Picker
-
Failing jobs (with the same error): Manually Curate Exposures, Manual Picker, Extract From Micrographs
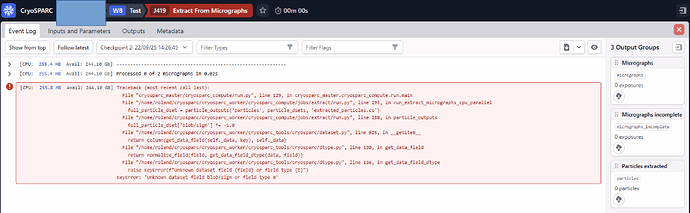
The error messages point to a binfactor of 0 (e.g. ZeroDivisionError: integer division or modulo by zero). It seems the bin factor is defaulting to 0 — is there a way to force it to default to 1?
Additionally, in the metadata I see:
"micrograph_blob/is_background_subtracted": 0
But in reality, my micrographs are already background-subtracted. Is there a way to modify this field so CryoSPARC recognizes it correctly?
Any guidance on whether this workflow is supported, and how to properly adjust the metadata/binfactor, would be greatly appreciated.
Best,
Hao
> Sep 20, 2025, 12:29:09 AM
Traceback (most recent call last): File “cli/run.py”, line 105, in cli.run.run_job File “cli/run.py”, line 210, in cli.run.run_job_function File “compute/jobs/extract/run.py”, line 208, in compute.jobs.extract.run.run_extract_micrographs_cpu_parallel File “/home/cryosparc2/cryosparc/cryosparc_worker/compute/jobs/extract/extraction_cpu_parallel.py”, line 122, in do_extract_particles_single_mic_cpu mic_bin = mg.bin_mic(mic, bg_binfactor) # slow.. ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^ File “/home/cryosparc2/cryosparc/cryosparc_worker/compute/micrographs.py”, line 606, in bin_mic clipx = nx // binfactor ~~~^^~~~~~~~~~~ ZeroDivisionError: integer division or modulo by zero