trying to make a mask for signal subtraction, would be nice to get the inverse mask (volume tools).
I do a refine, make a mask of the region to keep, then import that. Open volume tools.
volume – load the output map from the refine above
mask – load the “keep mask” from above
invert mask – click button
type of input – map (have also tried mask)
type of output --mask
every time I get a blank output. Can’t find tutorials for how to do this and I’m clearly too dumb.
thanks.
It’s a bug IMO (@apunjani?) . Uncheck the option “fill holes”. When inverse mask is selected, cryosparc treats the zeroed region “inside” the inverse mask boundary as a hole, leading to a completely blank volume (all ones).
Cheers
Oli
Hey guys, thanks for reporting, we’ll be looking into this shortly
thanks for the response. Did not fix the issue though. No worries I will make the mask another way. Thanks again.
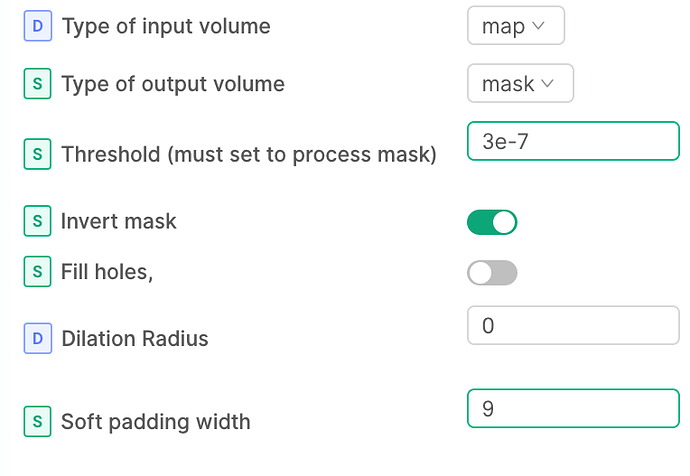
Huh that’s weird - to be clear, the parameters should look something like the attached. In my hands this correctly makes an inverse mask (assuming the threshold is set appropriately).
Cheers
Oli
thanks for the response. I was doing it wrong.
I provided the input volume and the mask of the area I wanted to subtract. I was hoping this would yield my initial volume minus the provided mask.
Misunderstanding of how this feature works.
What I did instead was provide the “keep” map (not mask) only and simply asked for an inverse mask for that map. It creates a massive cubic mask that will remove ALL signal around the region of interest when doing signal subtraction. I assumed this kind of solvent zeroing would be a bad thing and that I should be subtracting just the areas with protein rather than the entire box.
Anyway we’ll see how it pans out. thanks for the help. Hopefully this post helps someone in the future having a hard time getting the inverse mask tool to work.
Hi all,
In cryoSPARC v3.1.0, the “Invert mask” option should work with the “Fill holes” on — the new behaviour is that the input mask is first thresholded, then has holes filled, and then is finally inverted. This will generate a mask that covers the whole box except for the initial region. Note that as @orangeboomerang pointed out, this will cover the solvent too – we don’t test this workflow very often so perhaps you may get better results by instead using the volume eraser tool in Chimera, and manually creating the “inverse” mask (as detained in our mask generation guide page).
Best,
Michael