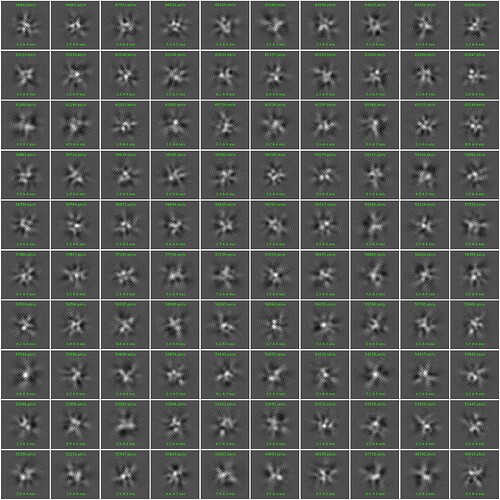
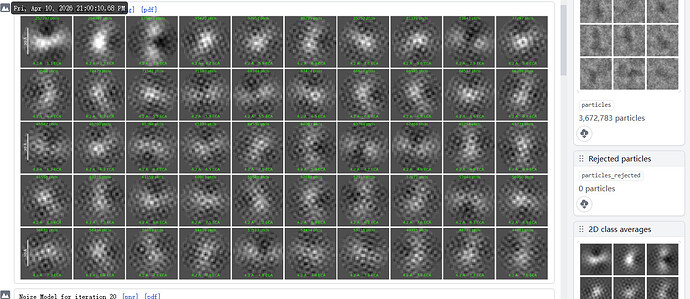
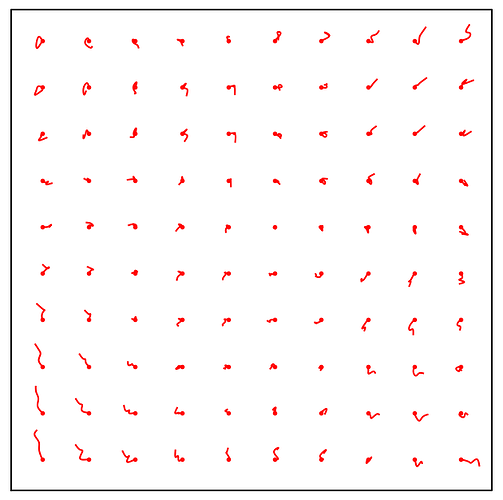
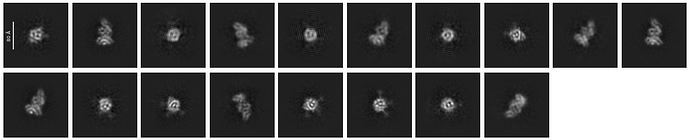
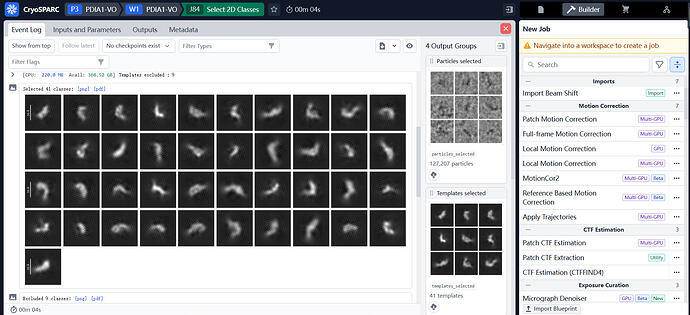
I am trying to generate 2D classes for my data and I have been getting these honeycomb like noise in my 2d classes. I have tried both blob pick as well as manual pick so far. Any inputs will be much appreciated on how to get rid of these noisy patterns.
Looks like overfitting.
What do the micrographs look like? The 2D looks like it may be a small protein at high concentration. Try imposing a mask rather than just relying on the windowing function. Or limiting resolution to perhaps 8Å. An example micrograph might indicate whether some careful processing can work, or whether you might need to go back and try with new grids (possibly at a lower concentration)…
@rbs_sci thanks for your reply. Indeed these are small protein oligomers which are stable at high concentration only. I have tried imposing circular mask diameter but it didn’t help much.
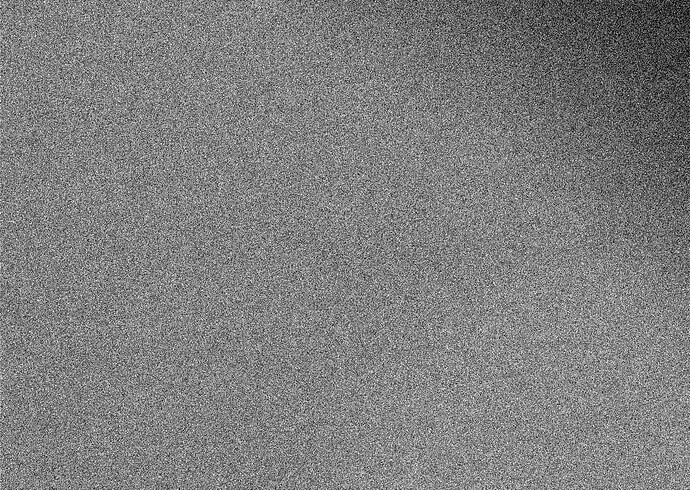
Here is the representative micrograph.
Thanks. OK, what do your picks look like? Very little jumps out at me on that micrograph, just in the top right and something in a band across the middle there are some features which may be particles, so if your extracted particles cover all of it, you may be including a large number of boxes which just contain vitreous ice (and noise) which may cause issues.
What defocus range are you working across?
Forgot to say earlier, but welcome to the forum. ![]()
Hi. I had the similar 2d classification before. In my case, it turned out to be the patch motion correction did not work for my movies. I changed to the full frame motion correction. Then everything went fine. You could take a try.
there are no clear particles here. I would recommend collecting some images at high defocus (~3-4um), where particles, if present, should be very obvious. Do other micrographs have clearer particles? What is the approximate molecular weight of your oligomer?
Hi @rbs_sci, I am trying to improve my 2d classes for a small protein+oligo complex. Just double-checking your comment here - when you say ‘imposing a mask’, you mean for the refinement jobs? Or did you mean a circular mask based on the particle dimensions?
I see my particle but 2D classification picks up a lot of noise on the bottom view classes of the particles even after tightening circular mask parameters. I have attached a picture here too. Thank you!
That’s what I meant, yes. ![]()
Your 2D looks quite promising. If running CryoSPARC 5, have you tried ab inition refinement (a new job type which holds gold-standard FSCs valid even during ab initio)?
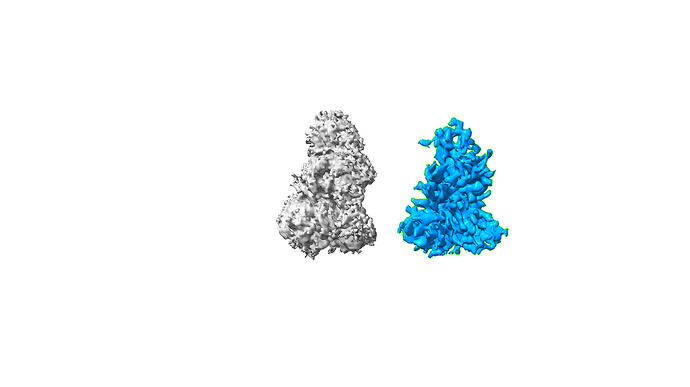
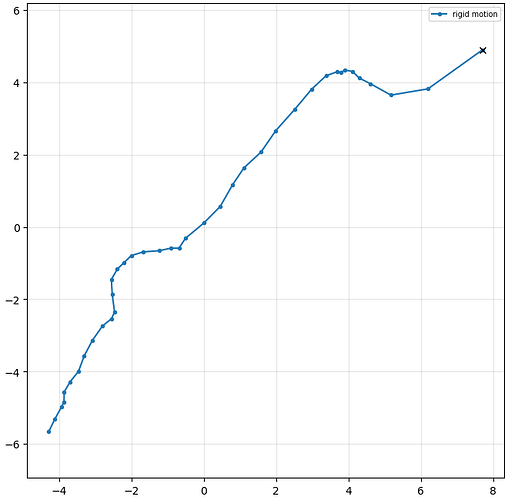
Thanks @rbs_sci ! I have been using V4.7, but will be updating to V5 soon. I followed @olibclarke 's HR-HAIR method and unfortunately, I am still finding it difficult to get good 3D refinements, due to overfitting and somewhat ‘streaky’ noise. I have attached my ab initio (grey) and homo refine (blue) below. Thank you!
Hi @rbs_sci, I have a similiar problem when I do 2D classes for a dimer protein(M.W. 55kD). I try to improve my 2d classes by repeatedly performing template picks and 2D classification. I changed NCC socre and box size for many times. However, I always get honycomb noise in 2d class. Do you have any idea for this problem?
I have attached a photo here too.Thank you!
Your box size is way too small - start by increasing it. i would suggest 200Å to start
If that doesn’t help, and you see clear, well defined particles on the micrographs, I would consider the possibility that your ice is too thick, or that something went wrong with the scope during collection
Oli is absolutely right - the box is far too tight. 150-200 Å, play with mask diameter as necessary.
Thank you oli. You are right. I changed the box size and the result was much better.However, there were still a lot of honycomb noise in 2D class.So why is that?
I have attached a photo here.Thank you very much!
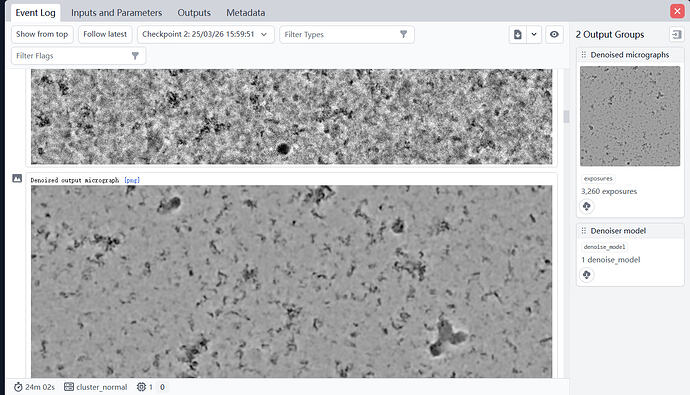
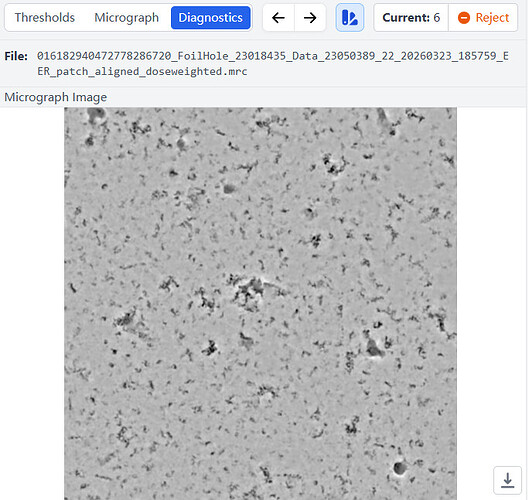
I upload a denoised picture. Most of photos were similar like this one.Do you mean my sample have a lot of ice contamination?
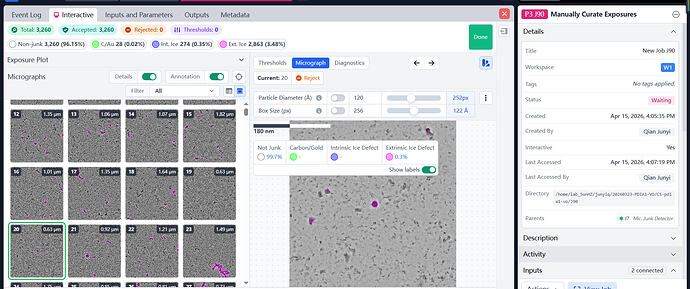
I used “manually curate exposures” to reject the junk micrographs. From your experience, could I use these data to solve a 3 angstroms stucture(pixel size:0.9519)?
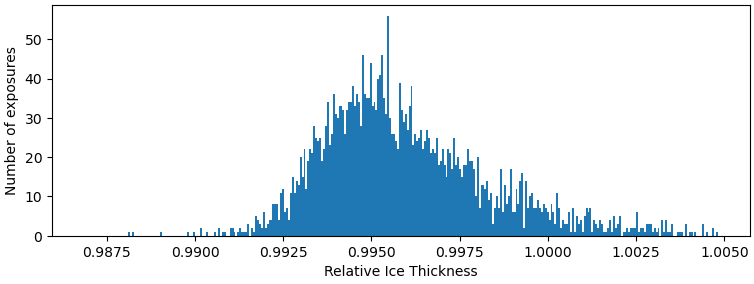
No, I mean maybe the ice in which the particles are embedded is too thick, which lowers the signal to noise ratio. What are the relative ice thickness values like? What does the power spectrum (FFT) of an example micrograph look like?
Sorry,I don’t know where to find the power sprctrum(FFT).But I attched a photo about relative ice thickness from ““manually curate exposures” .
You can view the FFT multiple different ways, but the easiest is probably looking at the diagnostics tab in Curate Exposures. The presence of a strong water ring would indicate that your ice may be a too thick for small particles. These relative ice thickness values look okay, but I would always prefer to see the raw data.
the other possibility is that your particle is just too flexible/disordered/heterogeneous to give good 2D classes
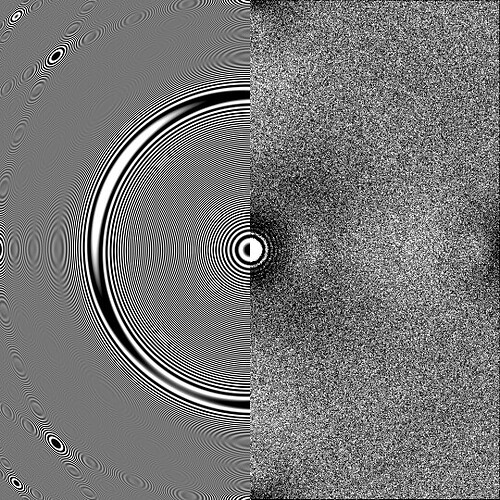
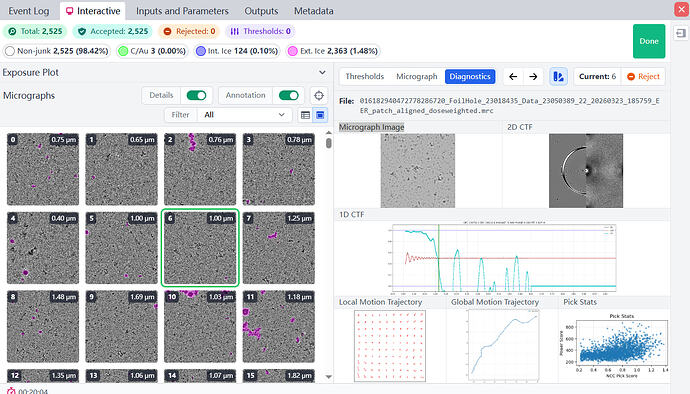
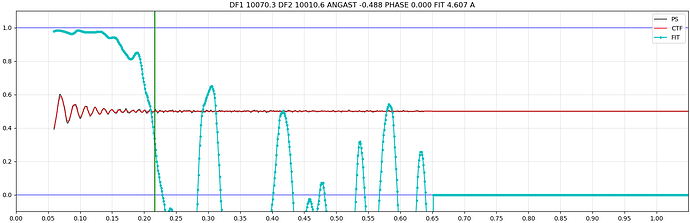
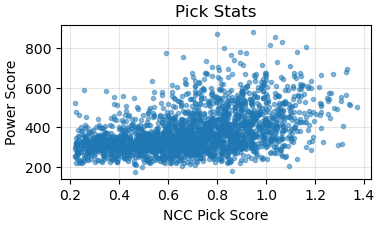
I selected a typical micrograph. Then I uploaded the Micrograph Image, 2D CTF, 1D CTF, and other diagnostic data for this micrograph. I would like to ask how to use this data to determine if the particle ice layer thickness you mentioned is appropriate, and what the quality of the raw data is?