Hi, community!
I’m new to this field, trying to export selected particles into image files, and I’m currently following the steps of this topic.
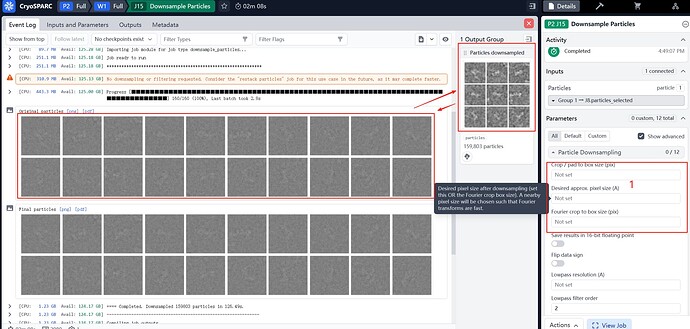
I created a downsample job and successfully obtained the mrc files.
Then I (LLM) write a python script (code is at the end) to transform mrc files to png files. But the png file seems full of nosie, does not looks like the particle showed in the cryosparc (see below).
And the next image is my 216*216 pixel PNG particle file.
Is this what my particles are supposed to look like? Any insights would be appreciated!
Code:
#!/usr/bin/env python3
"""
usage:
python mrc2png.py particles.mrcs --out_dir png_particles
python mrc2png.py *.mrc --threads 8
dependency:
pip install mrcfile numpy matplotlib tqdm
"""
import argparse, os, sys, glob
from pathlib import Path
import numpy as np
import mrcfile
from matplotlib import pyplot as plt
from tqdm import tqdm
from multiprocessing import Pool, cpu_count
# --------------------------------------------------
def norm_to_uint8(img):
img = img.astype(np.float32)
img -= img.min()
img /= img.max() + 1e-8
img *= 255
return img.astype(np.uint8)
def save_png(arr, png_path):
plt.imsave(png_path, arr, cmap='gray', vmin=0, vmax=255)
def process_one_stack(mrc_path, out_dir, prefix="particle"):
os.makedirs(out_dir, exist_ok=True)
with mrcfile.open(mrc_path, permissive=True) as mrc:
stack = mrc.data
if stack.ndim == 2:
stack = stack[np.newaxis]
base_name = Path(mrc_path).stem
for idx, img in enumerate(tqdm(stack, desc=base_name, leave=False)):
png_name = f"{prefix}_{base_name}_{idx:06d}.png"
png_path = os.path.join(out_dir, png_name)
save_png(norm_to_uint8(img), png_path)
def main():
parser = argparse.ArgumentParser(description="Convert MRC particle stacks to individual PNGs")
parser.add_argument("input", nargs="+", help="Input .mrc/.mrcs file(s) or wildcard")
parser.add_argument("-o", "--out_dir", default="png_particles",
help="Output directory for PNGs (default: png_particles)")
parser.add_argument("-t", "--threads", type=int, default=min(4, cpu_count()),
help="Parallel threads for multiple files (default: 4)")
args = parser.parse_args()
files = []
for pat in args.input:
files.extend(glob.glob(pat))
if not files:
sys.exit("No MRC files found!")
if args.threads > 1 and len(files) > 1:
with Pool(args.threads) as p:
p.starmap(process_one_stack, [(f, args.out_dir) for f in files])
else:
for f in files:
process_one_stack(f, args.out_dir)
print("All done! PNGs saved in", os.path.abspath(args.out_dir))
if __name__ == "__main__":
main()