I am having difficulty distinguishing real particles from carbon background and noise in my cryo-EM micrographs, leading to unreliable particle picking. I have already applied micrograph denoising and performed manual picking, but I still cannot confidently identify my complex (a 180 kDa trimer). Could this be due to poor micrograph quality or insufficient particle concentration? For context, I purified the complex by SEC, confirmed its integrity and purity by mass photometry after crosslinking, and applied it to the grid at 0.1 mg/mL.
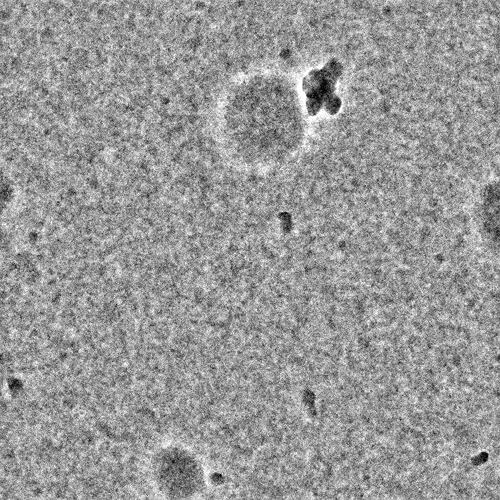
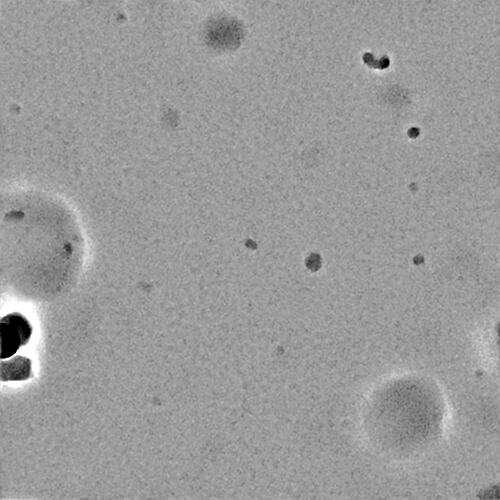
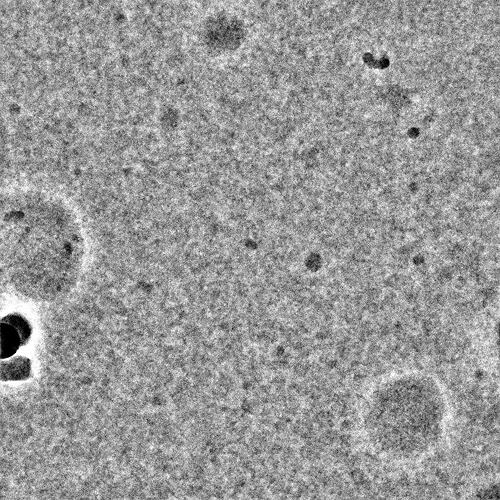
Can you post a sample micrograph (denoised & not denoised)? At 180kDa it should be obvious if it is there
Nothing obvious there I’m afraid - you may need higher protein concentration?
I agree that it is almost certainly a concentration issue based on your micrographs, which look fairly empty minus the junk. I usually tell my users to start at around 2 uM complex and go up and down from there – for a 180 kDa complex, 0.1 mg/mL is 0.5 uM, so you have a way to go. Unfortunately, cryo-EM concentrations do not scale linearly, so a 4x increase in concentration will not generate 4x the particles. Having too many particles is an easier problem to solve than too few, so I often encourage for starting to try to screen up to 8-10 uM if possible in addition to other concentrations. Detergents also lower apparent concentration and will require you to bump up concentration even more. Good luck!
Edited to add: if your protein absolutely can’t get up any higher, you can try doubling the sample application (apply protein, blot, apply more protein, blot again, plunge). Blotting is a process that concentrates sample, so you will end up with more sample on the grid than if you apply and blot once.