Yup, that’s what I meant. Mine went from looking like this
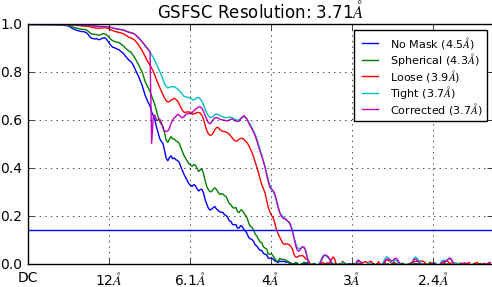
to this
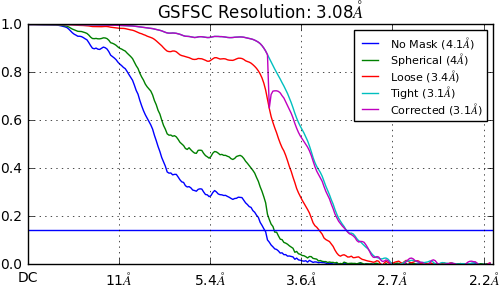 .
.
Of course, I was also imposing symmetry in the first one and that can be a big factor. Also, if you have a lot of overlapping boxes, you tend to see this kind of behavior (i.e. a bump in the FSC at a lower res.)
A better explanation is here