Hi all,
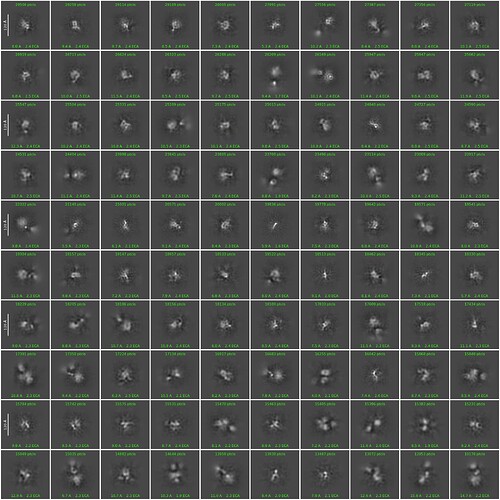
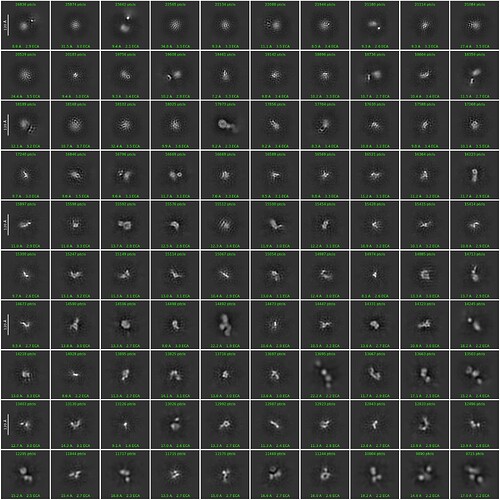
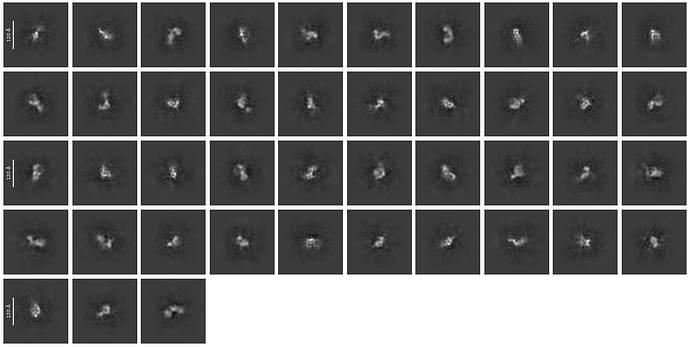
I am trying to get decent 2d classes for a ~130kDa protein. I tried to start from a template picking using a template generated by Alphafold-predicted structure. I attached the best class picture so far here. This is coming from the 2d-class job with customized parameters (50 classes, 40 Online-EM iteration, 400 batch size with force max on).
What I know is that this protein is around 120A in diameter, and according to alphafold structure the domain-domain linkage is quite flexible (so maybe 120A estimation is kind of exaggrated). I can see some features in the classes and selected ones look promising, but I think the signal in each class is still blurry and messy. Do I need to further refine the 2D classes before through them into Ab-initio? Thanks a lot!
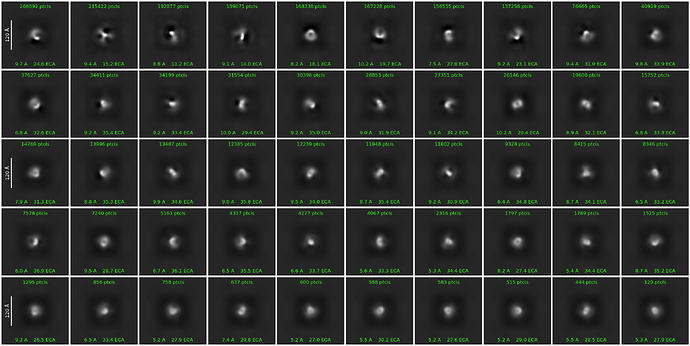
Btw, I also tried to switch off the force max pose option with additional changes (force max off, 80 o-em iterations, 400 batch size) and I got this.
Honestly it’s my first time to try force max off and maybe I should increase the full iteration numbers according to previous discussions. FYI, currently I am using a box size of 324 pixel with the pixel size of 0.868A/pixel.
1 Like
Hi @Chengtao ,
Some thoughts here:
(1) I usually advise my users against template-based picking from an alphafold structure. There have been numerous discussions on the forum about this, and generally it is probably OK if you lowpass filter your imported alphafold map to around 20A before you use it as a template, but there is always the danger that you are biasing your picking or generating Einstein-from-noise if you do this. This is especially true if your protein is flexible and the components are not necessarily oriented towards each other in the same way as the alphafold has predicted. In my opinion the better strategy is almost always to use a blob picker and then use templates generated from blob picking to do template-based picking if needed. I do encourage folks to use the template generation job to get an idea of what their 2Ds could look like, but don’t encourage them to use those for template-based picking, especially on a new project. If blob picking can’t pull something out of your data set, it usually means it isn’t a good data set to start with and would make me even more suspicious that your templates are pulling out noise.
(2) Have you completed multiple rounds of 2D classification and selection? It looks like there is a lot of junk in your classes and you do almost always have to do multiple rounds of cleaning in order to see “good” 2D classes with tricky proteins. Especially in your second post it looks like you have some good 2Ds in there among the noise! You seem to have lots of particles to start with, so even with multiple rounds I think you will have sufficient particles to play around with.
(3) We use a smaller maximum alignment res and resolution in 2D classification with small particles to force a larger internal box size to be used. I think it helps, we got it from somewhere on the forum. We weren’t finding changing the online-EM iterations to improve our small particle 2Ds, so we stopped using that, but do use force max over poses/shifts false and 400 batchsize. It is slow but does give better results for proteins <150 kDa or so.

(4) It is always worth trying more extensive classification/cleanup in 3D instead of in 2D if you don’t feel like you are getting good separation of classes in 2D, but I would still get rid of the classes that are obviously just noise/junk first. CryoSPARC is fast enough that it is worth trying both and seeing what gives you the best results 
Good luck!
2 Likes
Hi tlevitz,
Thanks for your advice!
(1) We tried blob-picking parallel to the Alphafold prediction-based template picking. The 2D class look similar to the template-based. I would try different parameters in blob picking and see whether iterative classification-picking will generate a good particle set.
(2) This is two round of 2D classification, and I am planning to do more rounds. Thanks for your advice!
(3) Thanks for your suggested parameters! I am sticking on O-EM iteration and batch size parameters. I haven’t tried to change the alignment res yet and it looks definitely worth a try!
(4) I did an ab-initio refinement followed by a heterogenous refinement. All 8 classes looked no obvious details and a lot of noise signals are still remained. So I think I may need to push more on 2D so far.
By the way, I have an additional question. If the 2D classification results are very blurry, does that imply that the subsequent 3D reconstruction steps (such as ab-initio reconstruction and heterogeneous refinement) will also perform poorly, resulting in low-resolution maps with blurred density and significant background noise?
Big thanks for your help!
Chengtao
Without seeing all of your processing, here is what I would suggest:
-I would try to be pretty stringent with your selection of 2D classes at first, because many of these seem to be junk / noise (particularly in the second image). Usually you don’t have to do tons of rounds of 2D, but you can try for example two workflows, one where you are very picky about choosing classes where you only pick the best classes and throw out most of your particles (or “particles”), and one where you are looser and try to pick anything that is generally OK, and then proceed with processing both side-by-side and get a better idea of what things look like from there. Also, if alphafold templates look similar to your blob picker templates, definitely use the blob picker templates and then you can assure yourself you haven’t introduced any bias  You can use good classes from the blob picker and input those into template-based picking if you still want to do template-based picking.
You can use good classes from the blob picker and input those into template-based picking if you still want to do template-based picking.
-Did your ab-initio refinement look like what you expected it should look like? I would worry that you have too much junk in there and so are aligning on junk. Additionally, what did you use as the 8 inputs for the heterogeneous refinement and why 8 classes? Generally, I don’t advise people to do that many classes in heterogeneous refinement almost ever, unless you genuinely expect to have 6 or so different species / distinct conformations in there – you can do 3D classification to see small differences later if you need. What I tell my users to do, at least for a new project, is to make 1 ab-initio for each species / very distinct conformation present or expected, and then do 1-3 noise models as well (credit to @olibclarke for this, I am pretty sure). To make noise models you can run the ab-initio and then stop it immediately when it starts showing a number of particles in the card view, and then mark the job as finished (You can also run an ab-initio job with the following parameters and let it go to completion, if you are like me and forgetful, and it seems to work pretty well in our experience – we just have a blueprint job with these parameters now:)
Number of Ab-Initio classes: 3
Num particles to use: 10
Number of final iterations: 1
Number of initial iterations: 1
Final minibatch size: 25
Initial minibatch size num iters: 1
Initial minibatch size: 25
Class similarity anneal start iter: 0
Class similarity anneal end iter: 1
Initial learning rate duration: 1
Maximum resolution (Angstroms): 25
Regarding your question about the blurry 2Ds, that can be due to a number of things. You mentioned your protein is flexible, so that is probably part of it! Protein flexibility and noise/junk particles are not necessarily related, so you can end up with a low resolution structure with very little noise / junk or a higher resolution structure (but probably not highest-resolution structure) with junk still in there. Sometimes blurry 2Ds indicate that you need to do more stringent classification, but often it has to do with something with your protein itself. And since most of the later steps do pose / alignment assignments from scratch, getting slightly better aligned 2Ds isn’t going to necessarily give you any leg up on the later steps (some of which may allow for un-doing motion to some extent, although the smaller your particle is the harder it will be – you may have to try some solutions on the sample / specimen preparation side of things).
Hope this helps clarify some things!
2 Likes
Hi tlevitz,
I am currently starting over from blob picking and see what happened. The latest 2d classification based on previous reply looks not convergent (max align res 3A max res 3A force max off batch size 400).
And I also ran a 2d classification with these parameters (max res 4A batch size 400 O-EM iterations 40 full iteration 3). The classes are very blurry.
It may be worth mentioning that all the 2d classifications are based on denoised micrograph after Micrograph Denoiser job. I think it may not be a good idea to start from that since the contrast is not good?
My ab-initio job (8 classes) results were bad. My previous experience on some big complexes relied on 6-class ab-initio to separate different conformations and junks, but I agree with you that I should try less classes to make a consensus model first, despite the flexibility. After several iterations of 2d-template picking, I will try your parameters listed above!
Also, I am curious how to use the noise model you mentioned above:)
Thanks for your detailed reply and suggestions!!!
Chengtao
Here is the 2d class image for the blurry 2d
Hi @Chengtao,
Are you using an inspect particle picks job when you use the blob picker? I ask because it seems like you may have far more particles in the blob picking job than in the template-based picking. When particle picking with any (blob or template-based) picker, you should always inspect how the job is doing visually on the micrographs and adjust picking parameters accordingly. It’s OK to have some junk, but you want to be relatively sure that you aren’t drastically overpicking or underpicking your particles. You can always try different picking parameters on a small subset of micrographs and that will run very quickly, and then the parameters you like you can run on the entire data set. If you are confident that you have extracted mostly true particles, you can always try using fewer 2D classes in the initial rounds to really just separate junk from things that could be a true particle. Sometimes I only use 10 or 15 classes in an initial round.
re: denoised micrographs, I generally don’t use them, but if anything they should improve your contrast (and you should be extracting from the original micrographs anyways, so the 2Ds will use particles from the not denoised micrographs). Take a look at how picking works on both with inspect particle picks and then decide visually which one looks like it is doing a better job. If you can’t see your particles visually without denoising the micrographs, that is often a red flag that your data quality is poor, but denoising can help grab particles that are otherwise missed.
For noise models: once they are generated, you can attach them as initial volumes along with your (usually 1 or 2) “true” ab-initio models, Then, those noise models will hopefully grab junk particles so that your true models are only refined with true particles and you can use those particle outputs for further homogeneous (or local or nonuniform) refinement(s).
And just a note / reminder that you likely can’t force a good structure out of a extremely small, flexible, and low contrast data set. While it is absolutely critical to try different processing techniques and optimizing your picking and classification parameters to make sure you are actually picking and classifying true particles from the micrographs, there is always to possibility that the sample / specimen is what needs to be optimized and no amount of processing on the back end will change that.
Hi @tlevitz ,
I used inspect picked particles job to verify picked particles and remove the false-positive junks (which should have very high power score?).
I am currently using the original images and start over to do these picking jobs. Also, I really appreciate the explanation on the noise model and all the advice above! I can try to generate one in the ab-initio job!
Yes, I think this grid is suboptimal. The protein expression is good but we do find some troubles in entering the holes and uneven ice thickness. I may try some detergent or different grid type to figure it out.
Thanks a lot,
Chengtao
1 Like