Hi,
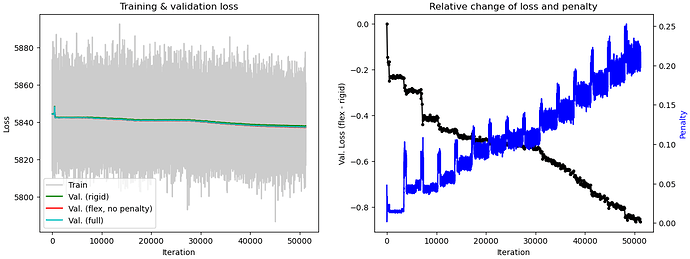
I ran 3D Flex with default parameters except for using 32 layers. I used 32 layers because my protein is relatively small (~250 kDa). I have very different plots from the tutorial video. What parameters should be first tuned to get better training results?
Thank you,
Joon
1 Like
I am having the same issue with the top left graph- everything is lumped into one line and I don’t know why. It made me worry there would be no results, but I actually did get some decent results.
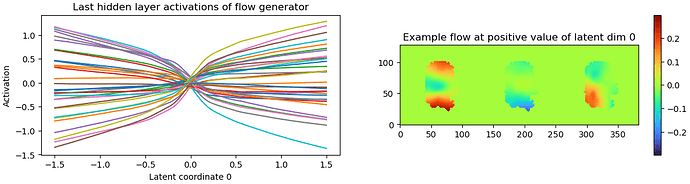
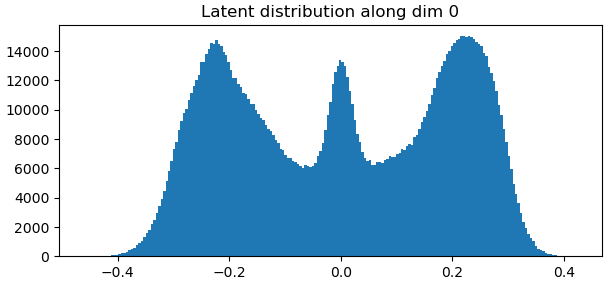
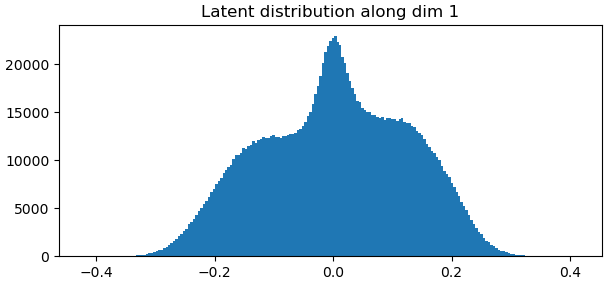
It looks like you may need to play around with some of the parameters such as the noise injection standard deviation and the latent centering strength to fix the bottom left “flow generator” graph. Maybe relaxing the rigidity might help as well. Once all of that is optimized, the “Relative change of loss and penalty” graph might improve.
1 Like
Hi! @jenchem I would like to ask which kind of machine are you running 3d flex refinement on. As the issue (3D Flex pytorch issue on RTX4090 - #13 by wtempel) exists, I can not run 3d flex in cryosparc v4.2.1 on RTX 4090. I have confirmed that cryosparcw call nvcc --version returns 11.8. But in CryoSPARC v4.2.1, after running “/home/xx/cryosparc/cryosparc_worker/bin/cryosparcw install-3dflex” to install pytorch with CUDA version 11.8, it returns the “1.13.1+cu117”.
Our GPUs are NVIDIA GeForce GTX 1080, and CUDA version is 12.2.
I know I’ve run 3D Flex beta on an earlier cryosparc v 4.2 and can confirm that it also works with this setup running v 4.3.1.
I hope you can find something that works!
Thanks for your reply! And did NVIDIA GeForce GTX 1080 (8GB GPU RAM) report “RAM not available”? I tried NVIDIA GeForce GTX 3070 (8GB GPU RAM) and 32GB RAM, but it reported this error. So I am finding a machine that can satisfy RAM requirements (3DFlex train job - RAM unavailable - #2 by apunjani) and not GTX 4090 at the same time.
I am working with a very large molecule but have been using the 128 box size for training for this reason- it is true that it’s quite a memory-intensive job.