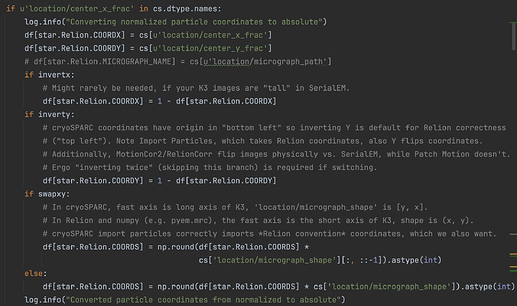
I am trying to revive this topic. My particles in Cryosparc through Patch Motion > Patch CTF > … > Refinement.
I can convert everything well according to the guideline from pyem using Methods 2. Extracted the particles and did a test reconstruction, and everything looked good. So particle coordinates & orientations are well converted by pyem. But when I polished it, the output particles were not at the right location at all. Does anyone see anything similar?
Extracted the particles from the cryoSPARC micrographs, right? You have probably needed to invert the coordinates and flip the CTF values.
Hi Daniel,
So apparently, to polish, I need to invertY and flipY (flip CTF as you said). But to test reconstruction by extract from the coordinate and reconstruct, I would use not InvertY (with --inverty option). It is a bit confusing but it works.
In this case it’s because in your “new” extraction test the particles are coming from the same drift-corrected micrographs you’ve had all along in cryoSPARC. Then only when you run polishing are you getting something based on Relion looking into the movies directly.
If you would instead run motion correction from scratch in Relion, then those new drift-corrected micrographs in Relion would have the same conventions as the Relion polishing job. (And in that case, extracting & refining would really tell you if Relion is getting the right coordinates. It would also let you use definitely correct CTF and alignment parameters, instead of the ones converted by me from whatever was estimated in cryoSPARC, based on flipping the images. Finally, Relion would also be able to start the polishing trajectories from the local patch trajectories found during initial motion correction. For that reason this is my recommended approach).
I’m running into the same issue as other users (“There is no movie metadata STAR file for any micrographs!”) – apparently, in a bid to consolidate data and save storage space I deleted the files for the initial “Import Movies” job, so while I have motion-corrected .mrc files from the subsequent “Motion Correction” job, the corrected_particles.star file points to un-corrected .tif files from the Import job that do not exist anymore.
When I re-run or clone the import job, the generated .tif files have different prefixes (file names formatted as {prefix}_moviename.tif) so the file names are not identical to the previous files. Is there any way to remedy this issue beyond full reprocessing from the initial data import?
@forrest If the problem is that the path to the tif files has changed and you did not delete or otherwise modify the original Import Movies job, you may be able to update the targets of the
symbolic links inside that original Import Movies job’s imported/ subdirectory (guide).
Hello Everyone,
I apologize for reopening the this thread again but I had a question in similar lines to this discussion. So if one polishes their particles and then re-extracts the polished particles in a binned box ( 2 times binning ) to further clean the data, does one then extracts the polished particles or the unpolished particles?
If they are the unpolished particles, is it possible to get the information that was stored during polishing into the newly cleaned dataset (to improve the resolution) without actually doing polishing again?
Hi,
Re-extraction sources the averaged micrographs, so these will be equivalent to unpolished particles. You can also simply downsample the polished particles for curation instead to get to a similar place. Alternatively, just use the polished particles as is, provided memory footprint isn’t too onerous?
In the case of re-extraction/downsampling, after identifying and discarding suboptimal particle images, you can perform an intersect routine (Particle Sets Tools) to fish out the particles of interest from the original polished set.
Cheers,
Yang
Hello Yang,
Thanks for clearing my doubt regarding this.
I will try this out.
Thanks a lot for the help.
To supplement @leetleyang’s correct answer, just think of polishing as “fancy extraction” - if you want new fancy particles (e.g. at a different box) you will always have to re-run it. Unfortunately there is no feature to re-use trajectory information to make it faster the second time.
Off on a bit of a tangent, if what you wish to do is simply repeat a polish job but with different box/resampling parameters, there is a hack-y way of reusing previous trajectories/B-factors and skip ahead to the frame recombination step:
CCPEM: Relion 3.1 – re-extracting polished particles from movies
Last did it in RELION-4, but I imagine it should work with RELION-5 as well.
Cheers,
Yang
Okay thanks a lot, I will give it a shot.
We typically do at least one round of Bayesian polishing in Relion on our final stack of particles. In the past, we have reprocessed in motioncor with the particle coordinates from cryosparc. This worked fine, it was just super slow. So, I wanted to give this a try again. I think it is working nicely, for one of my datasets, I am not seeing any improvement in FSC resolution (same) but the map is clearly improved. This will save us a lot of processing time, thank you @DanielAsarnow
Hi @DanielAsarnow following up on this. It does work perfectly and there is no difference in our maps between particles processed by skipping being extracted/refined MotionCorr micrographs vs. going through MC processing once we have the same stack. Thank you!
I will say though that there is one serious problem we have found, if you make the star file with –inverty and –flipy as you recommend in the guide, the output particles are not right. –inverty gives the right coordinates.
–flipy does several things from what I can see:
rlnAngleRot = -180+AngleRot
rlnAngleTilt = 180-AngleTilt
rlnAnglePsi = -1*AnglePsi
rlnOriginYAngst = -1*OriginYAngst
rlnDefocusAngle = -1*DefocusAngle
None of these are right. As far as I can tell, only the defocus angle needs to be changed like so:
-180+DefocusAngle
Hope this helps! Let me know if you need me to help update csparc2star or to test.
@jcoleman thanks for the report. Could you clarify the two scenarios you described? Are your micrographs between CS and Relion upside down from each other when viewed in e2display.py or another viewer? What is your movie format, and were any “flip in [axis]” options ever used in cryoSPARC extract?
It’s possible to need –inverty only, also for only the CTF to need to be flipped if the values were from e.g. Patch CTF in CS w/ “flip in Y” extraction. So I suppose –flipy may need to become two arguments for the CTF and for the alignments.
@DanielAsarnow interesting, the resultant micrographs corrected in CS relative to Relion are not flipped though the coordinates are flipped in y without using –inverty. The movie format is EER, we never used any flip in axis options in CS extract.
We find the defocus angle needs to be flipped by -180 degrees also, not multiplied by -1.
One other thing that I noticed is that generated y trajectories are negative, compared with MC where they are positive. This does not appear to affect anything.
@Joleen the flipping issues are due to 1) a historical choice to flip all TIFF files in Y which is consistent in SerialEM/IMOD/EMAN2/Relion/WARP/UCSF MotionCor(X), and 2) cryoSPARC and Relion use opposite Y-axis origin conventions (top vs. bottom left) but cryoSPARC is designed to correctly import Relion-style coordinate files. Originally, cryoSPARC only did ab initio reconstruction and homogeneous refinement, so the default arguments for csparc2star.py reflect that case where the movies are corrected outside and particles are imported. Passing --inverty actually has opposite semantics, it sets the inverty condition below to False. I know it’s confusing but by the time Patch Motion and Relion Polishing (operating on movies, not micrographs) were available there were already a lot of people using the program in established pipelines and I didn’t want to suddenly break all of those.
EER files aren’t flipped by Relion, so it makes sense that you need --inverty.
However, since the images are not flipped, the CTF information and particle alignments should not need any modification. Rotating the defocus angle through 180˚ doesn’t do anything theoretically, because it’s the angle the major defocus axis (U and V, not X and Y) makes from the X axis and without high-order aberrations/coma we have Friedel symmetry. Multiplying by -1 OTOH makes sense if the image is flipped vertically, then the power spectrum is also flipped.
If there is an issue with the defocus angle values (without --flipy) then I think it has to be some changed implementation detail in Relion that demands angles within a certain range, which subtracting 180˚ happens to deliver.
Regarding the y-trajectories, I believe this is actually an issue. The trajectory code flips the trajectories because of the conventional difference to Relion. There’s no switch not to do this for EER movies. I suspect your data is good enough that polishing can align the particle frames even when the initial trajectories are bad. If you wouldn’t mind could you convert ~10 movies to TIFF with relion_convert_to_tiff[_mpi] and see if the trajectories are flipped or not vs. EER?
Hi @DanielAsarnow thanks! You’re right, there does not seem to be any difference in relion if the angle is rotated by 180 degrees.
I converted a few movies to tiff like you suggested, ran patch motion, and then converted them using cpsarc2star. These y-trajectories match more closely to the trajectories from MotionCor. So the y-trajectories are indeed flipped relative to EERs.
@jcoleman Thanks! I will put something in to deal with this (probably multiplying y-trajectories by -1 if the cryoSPARC imports were .eer - does that sound right?)
@DanielAsarnow that sounds good to me. Thank you!