hi devs
I have tons of high quality krios data collected on a two-component computationally designed de novo protein that has substantial movement engineered into it. In 2D averages both components are visible, however in 3D only one component is ever able to be resolved to side chain detail.
I attended Ali’s webinar on 3DVA and have read the tutorials. The output only ever shows blobs of density for the unresolvable component. It’s frustrating because it’s visible in 2D and so I suspect it must be some processing issue. I have tried every conceivable trick including subtracting signal for the resolved component but still cannot resolve the other component. I have also tried the various output options in the 3DVA display to no avail.
I have several designs where I’m running into this problem. I wonder if it is a good test case for 3DVA improvement? This is an extremely challenging problem and would love to hear any suggestions and/or if there is any time or interest to collaborate on such problems.
thanks
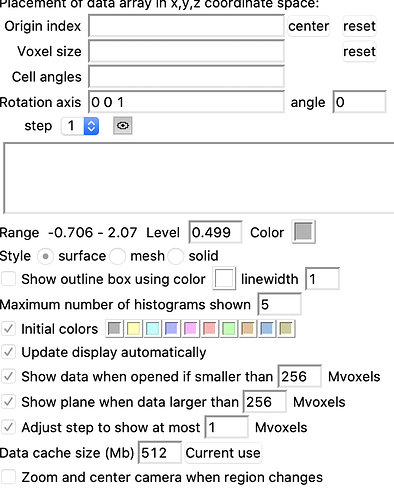
What do the actual 3DVA components look like? If you open those volumes and use Volume > Data display options > uncheck “cap high values at faces” and then look at their positive and negative iso-contours, are they capturing information at the expected resolution? How are you constructing the 3DVA mask in this case?
Yes, if you do not see the flexible part in 3D, that might be hard to make a mask that should include that region.
hi
thank you for the response.
As per the tutorial (or Ali’s webinar?) my mask includes portions that move a lot and portions that are stationary at about 50% of each. I have also tried a mask that includes only the part that is moving but the results are not helpful.
As for what the 3DVA components look like… I can share images if we take this to a private conversation, since it is a collaboration I don’t want to post here.
I visualize the outputs by “volume series” in chimera. I have tried the job with low pass to 10A because the expected motion is quite large. Also tried no lowpass. In either case I really dont ever see more than just a blob for the second unresolved component. Again it feels like there’s more detail in the 2D so I expect more in 3D.
I am also trying to high pass to 20A right now (along with lowpass to 10A). Trying again the “intermediates” output type after re-reading the tutorial which states this is the better option for samples with larger movmeents.
Often I see the one component is reasonably resolved
I dont see “cap high values at faces” in either chimera or chimeraX, see below.
Hi @orangeboomerang, thanks for your post - definitely an interesting project. I will reach out to you by email to discuss further.
thanks Ali I really look forward to it.